Perseids – Personalizing cancer therapy through integrated modeling and decision

Project name: Perseids – Personalizing cancer therapy through integrated modeling and decision
Research group: Intelligent Systems
Objective
Description
Perseids is a three-year multidisciplinary national project that involves several institutions such as IDMEC, INESC-ID, IT, FFCUL, IMM, CHLN (Lisbon, Portugal).
The goal of project PERSEIDS is to create accurate classifiers and to identify factors associated with disease outcome, which in turn will support the design of adaptive controllers and decision systems. The ultimate aim is to develop robust clinical decision support systems for personalized cancer therapy optimization by integrating patient multilevel data.
One of the main results so far that derives from its inter-disciplinary nature between engineering and medicine, which is to significantly strengthen the bridges between these two areas. The challenges being addressed so far include the dynamic modeling and control of physiological systems, machine learning and clinical data science tasks. We believe that this mathematical and computational approach, coupled with medical knowledge, will be the key to unravel organisms’ behavior and their responses to stimuli.
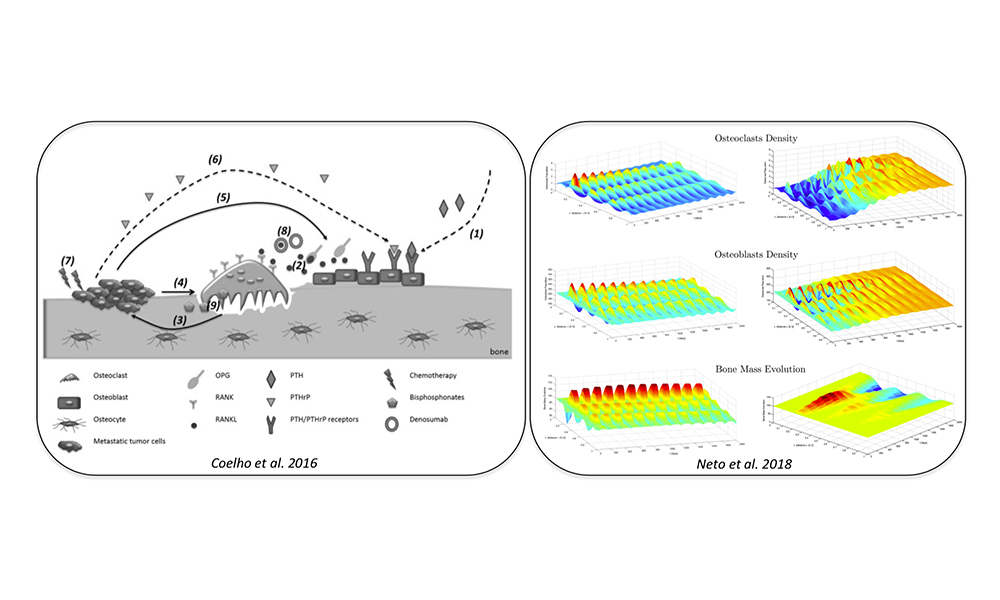
So far we have been progressing in modeling of bone physiology in pathological states, such as in cancer, by proposing novel models that including the pharmacokinetic/pharmacodynamics and the immune system responses. The application of sparse survival models has also provided interesting biomarkers for oncological diseases that can now be studied from a clinical point of view.

References
- MB Lopes, A Veríssimo, E Carrasquinha, S Casimiro, N Beerenwinkel, S Vinga (2018) Ensemble outlier detection and gene selection in triple-negative breast cancer data. BMC bioinformatics 19 (1), 168
- RM Coelho, JM Lemos, I Alho, D Valério, AR Ferreira, L Costa, S Vinga (2016) Dynamic modeling of bone metastasis, microenvironment and therapy: integrating parathyroid hormone (PTH) effect, anti-resorptive and anti-cancer therapy. Journal of theoretical biology 391, 1-12
- A Veríssimo, AL Oliveira, MF Sagot, S Vinga (2016) DegreeCox–a network-based regularization method for survival analysis. BMC bioinformatics 17 (16), 449
Contact Person
Susana Vinga
